Clonal and Functional Analysis Reveals the Capacity of Allograft T cells to Join the Circulating Pool after Human Intestinal Transplantation
Jianing Fu PhD1,2, Julien Zuber MD, PhD1,3, Brittany Shonts BA1,2, Kevin Bi4, Kathleen Yates4, Wenji Ma PhD5, Aleksandar Obradovic BA1, Suxiao Yang MS1,2, Siu-hong Ho PhD1,2, Laurence Turka MD6, Donna Farber PhD1,7,8, Nicholas Haining MD4, Yufeng Shen PhD5, Prakash Satwani MD9, Mercedes Martinez MD9, Tomoaki Kato MD7, Megan Sykes MD1,2,7,8.
1Columbia Center for Translational Immunology, Columbia University, New York, NY, United States; 2Department of Medicine, Columbia University, New York, NY, United States; 3Department of Kidney Transplantation, Necker Hospital, Paris, France; 4Department of Pediatric Oncology, Dana-Farber Cancer Institute, Boston, MA, United States; 5Center for Computational Biology and Bioinformatics, Department of Systems Biology, Columbia University, New York, NY, United States; 6Department of Surgery, Massachusetts General Hospital, Boston, MA, United States; 7Department of Surgery, Columbia University, New York, NY, United States; 8Department of Microbiology & Immunology, Columbia University, New York, NY, United States; 9Department of Pediatrics, Columbia University, New York, NY, United States
Resident memory T cells (Trm) are non-recirculating cells that persist in tissues. It has so far not been possible to assess whether migration between tissues and the circulating pool occurs in humans. Surveillance endoscopic biopsies on intestinal transplant patients permitted longitudinal and spatial analysis of relationships between intestinal T cells in allografts, peripheral blood and bone marrow (BM) using FCM, MLR, TCRβ DNA-seq and single-cell RNA-seq. Our published protocol was used to identify graft-versus-host (GVH) clones from pre-Tx donor lymphoid tissues1.
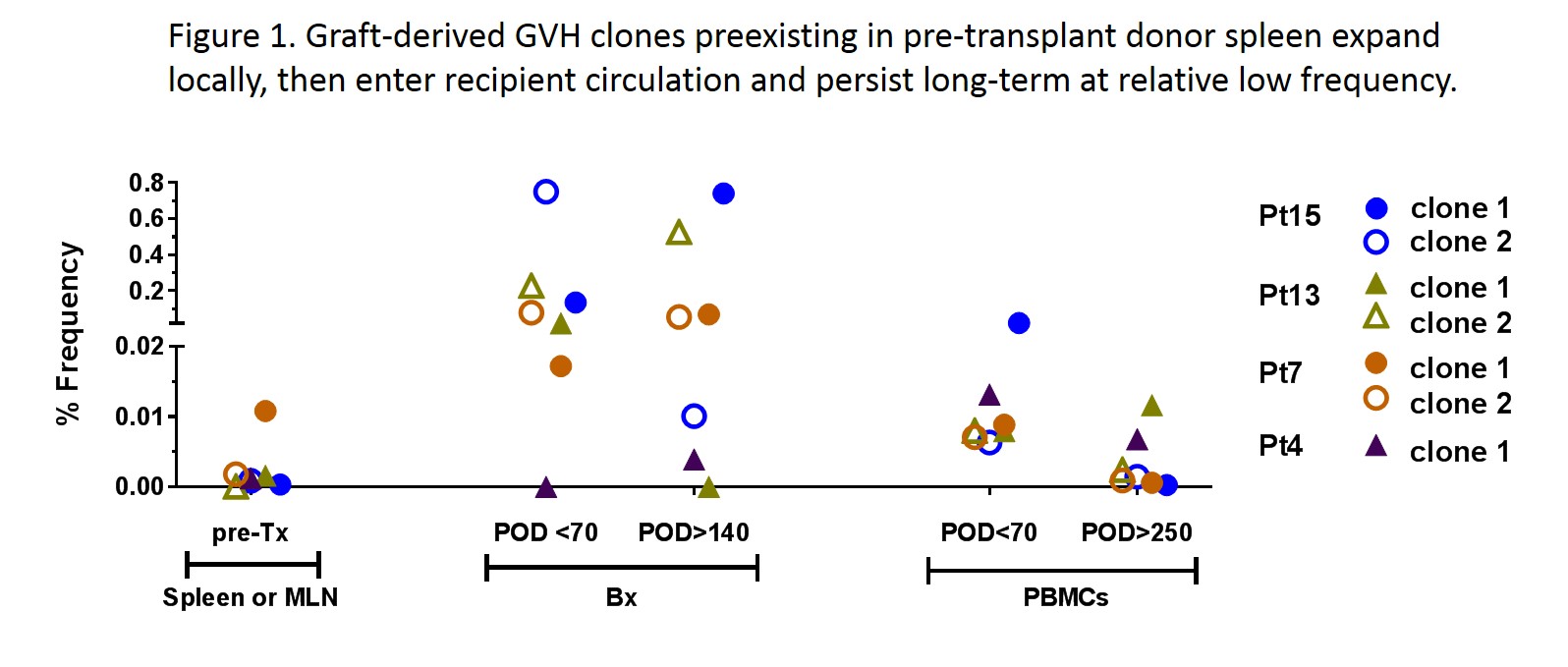
While the circulating donor T cells expressed a “blood” phenotype (CD28+CD69-CD103-), donor ileal lymphocytes in the graft displayed a Trm phenotype (CD28-CD69+CD103+/-). Donor T cells in post-Tx biopsies were greatly enriched for clones detected in both donor spleen and mesenteric lymph nodes (MLN), suggesting that Trm, including GVH clones, transferred in the allograft had a circulating memory counterpart. Clonal diversity analysis suggested that clones detected in both donor spleen and MLN were memory rather than naïve cells1. GVH clones identified in pre-Tx MLR by their CDR3 sequences persisted long-term in both the allograft (at high frequency1) and recipient peripheral blood (at low frequency) (Fig.1). These data suggest that clonal expansion occurred, likely in response to the early entry of recipient myeloid APCs into the graft1, and these expanded GVH Trm emigrated to the recipient circulation and acquired the “blood” phenotype.
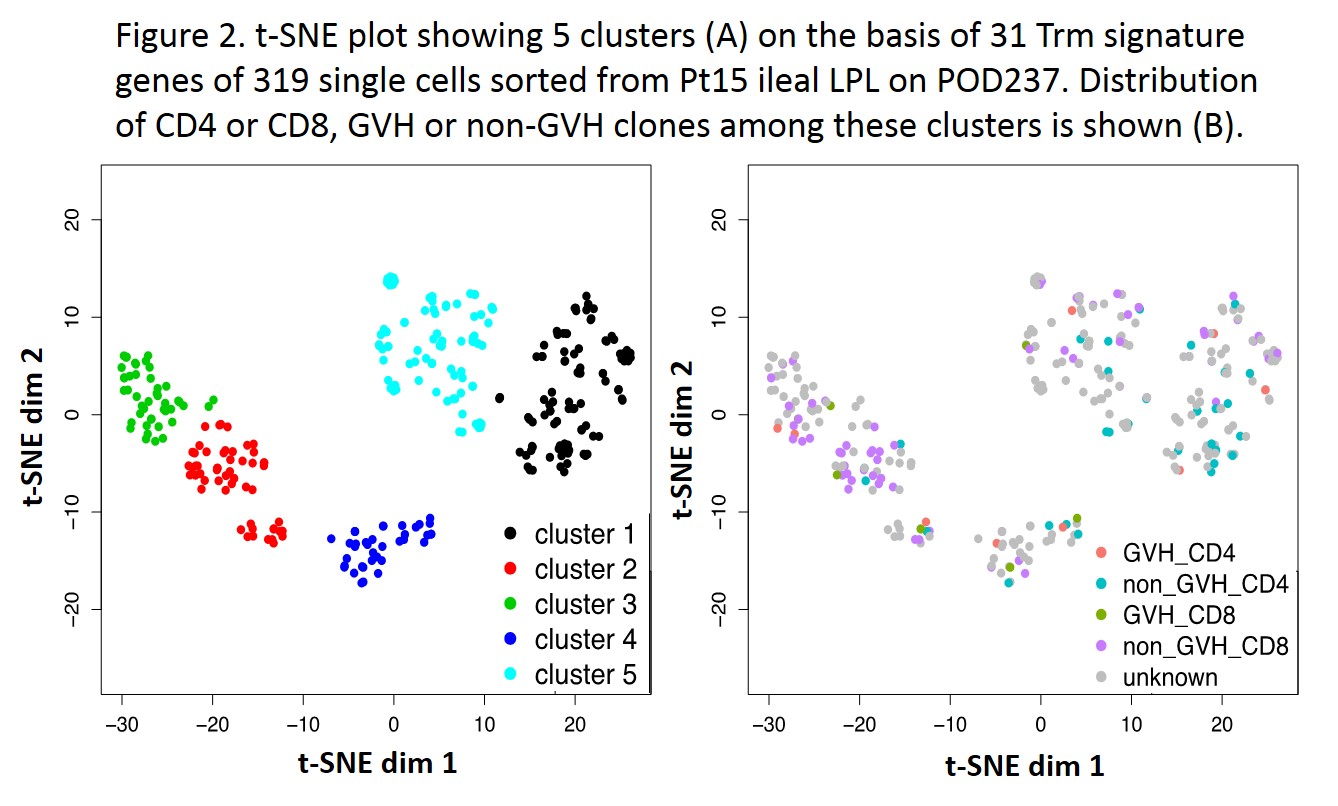
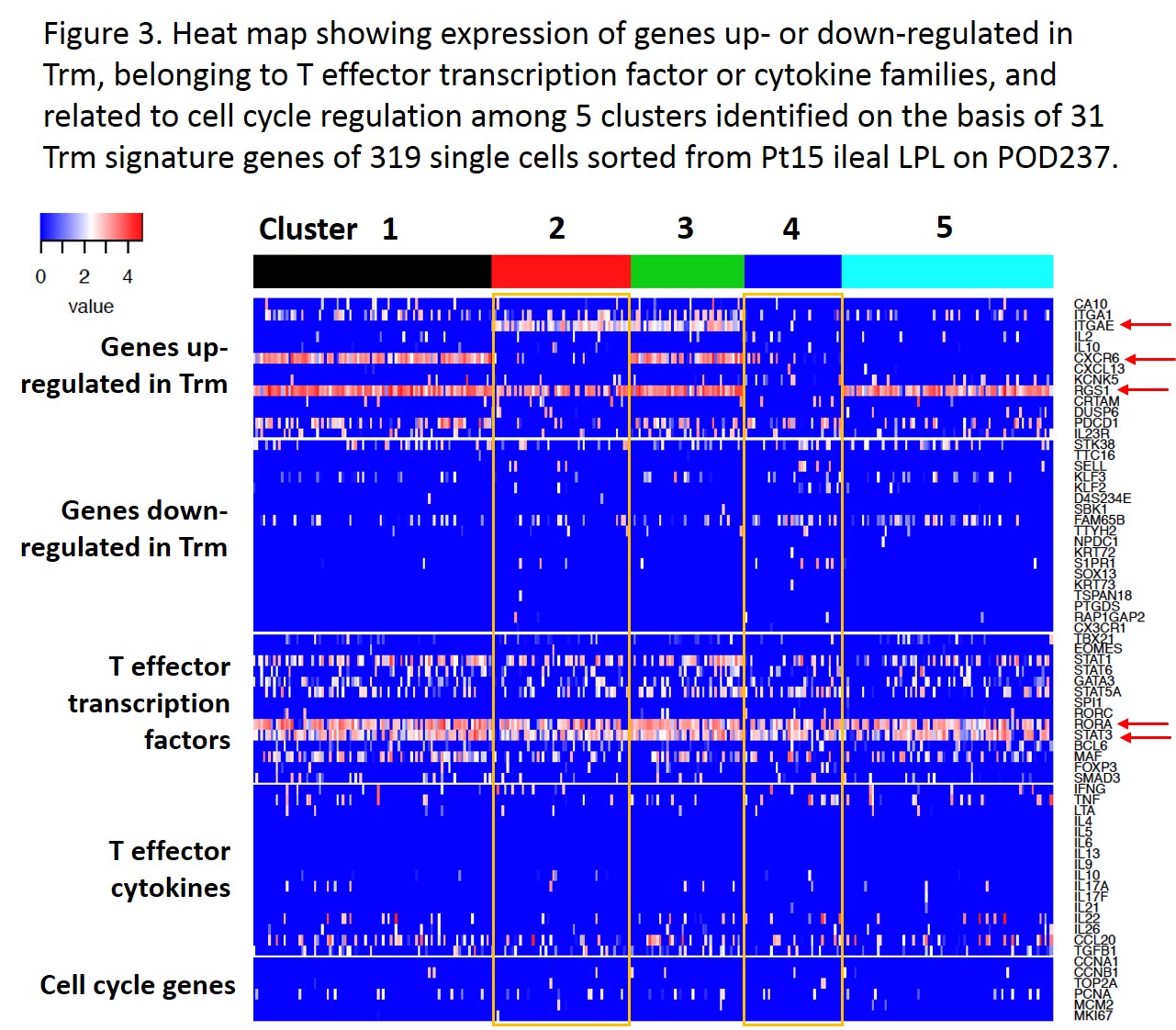
Single-cell RNA-seq data on GVH and donor-mappable non-GVH clones identifiable by their TCRβ sequences plus unknown TCR clones from a quiescent Day 237 allograft biopsy were divided into 5 clusters in t-SNE plots based on 31 Trm signature genes2 (Fig.2). All 4 “donor-mappable” clones (GVH or non-GVH CD4 or CD8) distributed into multiple clusters, showing no functional distinction between GVH- and non-GVH-reactive donor T cells in the graft. 4 of 5 clusters upregulated RGS1, a Trm retention signal, while other features of Trm such as ITGAE and CXCR6 were limited to certain clusters (Fig.3). Effector molecules including RORA and STAT3 were associated with several clusters.
Strikingly, in one multivisceral Tx patient analyzed, donor-HLA+ cells (0.04%) were detected in recipient BM on Day 105. Greater than 70% of these were T cells that were enriched for GVH clones among donor-mappable CD4s.
The temporal and spatial tracking of donor T cells and GVH clones derived from memory cells carried in intestinal allografts provides new insight into their migration patterns. Donor-derived Trm may leave the tissue of residence (intestinal graft) and join the circulating T cell pool, possibly after differentiating into effector T cells upon encounter with recipient alloantigen. Some of these cells may eventually reside in the BM niche.
1. Zuber et al., 2016 Science Immunology
2. Kumar et al., 2017 Cell Reports
The study was funded by the National Institute of Allergy and Infectious Diseases grant P01 AI106697. Research reported in this study was performed in the CCTI Flow Cytometry Core, supported in part by the Office of the Director, National Institutes of Health under awards S10RR027050 and S10OD020056. . Single cell RNA-seq study was supported in part by the Immune Tolerance Network supplement grant..
